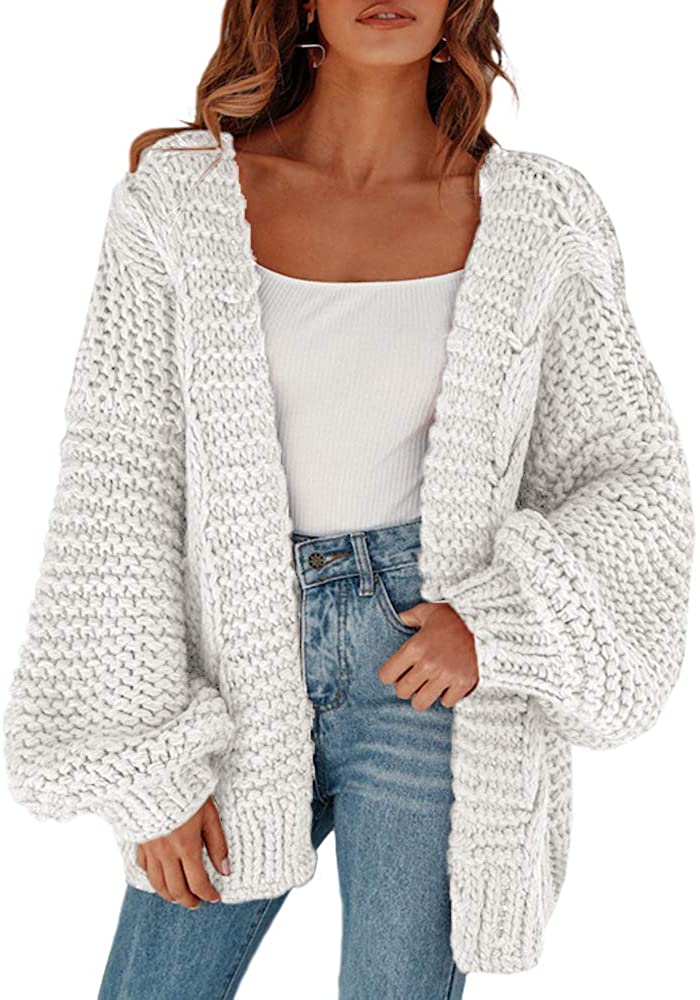
Micro-C XL: assaying chromosome conformation from the nucleosome
4.9 (776) · $ 17.99 · In stock
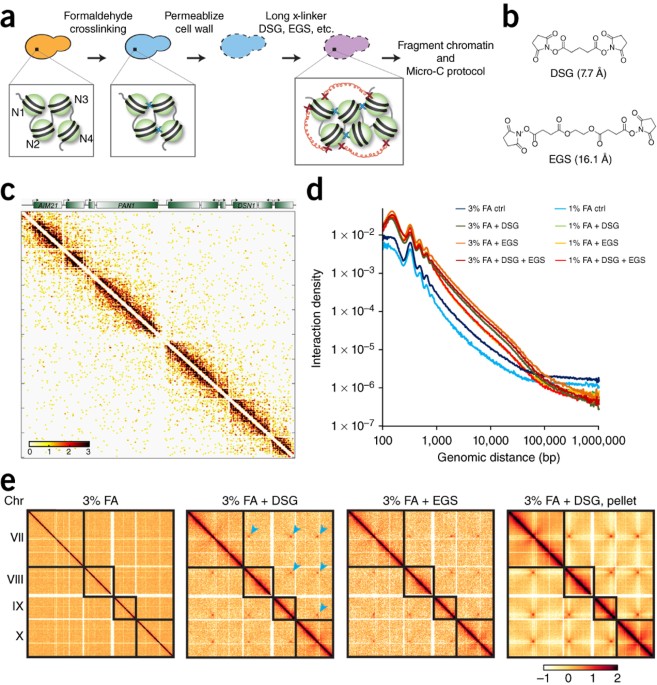

Cohesin residency determines chromatin loop patterns
![PDF] Mapping Nucleosome Resolution Chromosome Folding in Yeast by Micro-C](https://d3i71xaburhd42.cloudfront.net/65248ca1a2333ef2c553252fbc296a2b171e1812/8-Figure5-1.png)
PDF] Mapping Nucleosome Resolution Chromosome Folding in Yeast by Micro-C

3C and 3C-based techniques: the powerful tools for spatial genome organization deciphering, Molecular Cytogenetics

Novel biological insights revealed from the investigation of multiscale genome architecture - Computational and Structural Biotechnology Journal

Sub-nucleosomal Genome Structure Reveals Distinct Nucleosome Folding Motifs - ScienceDirect

Micro-C XL: assaying chromosome conformation from the nucleosome to the entire genome

Nucleosome positions alone determine micro-domains in yeast chromosomes
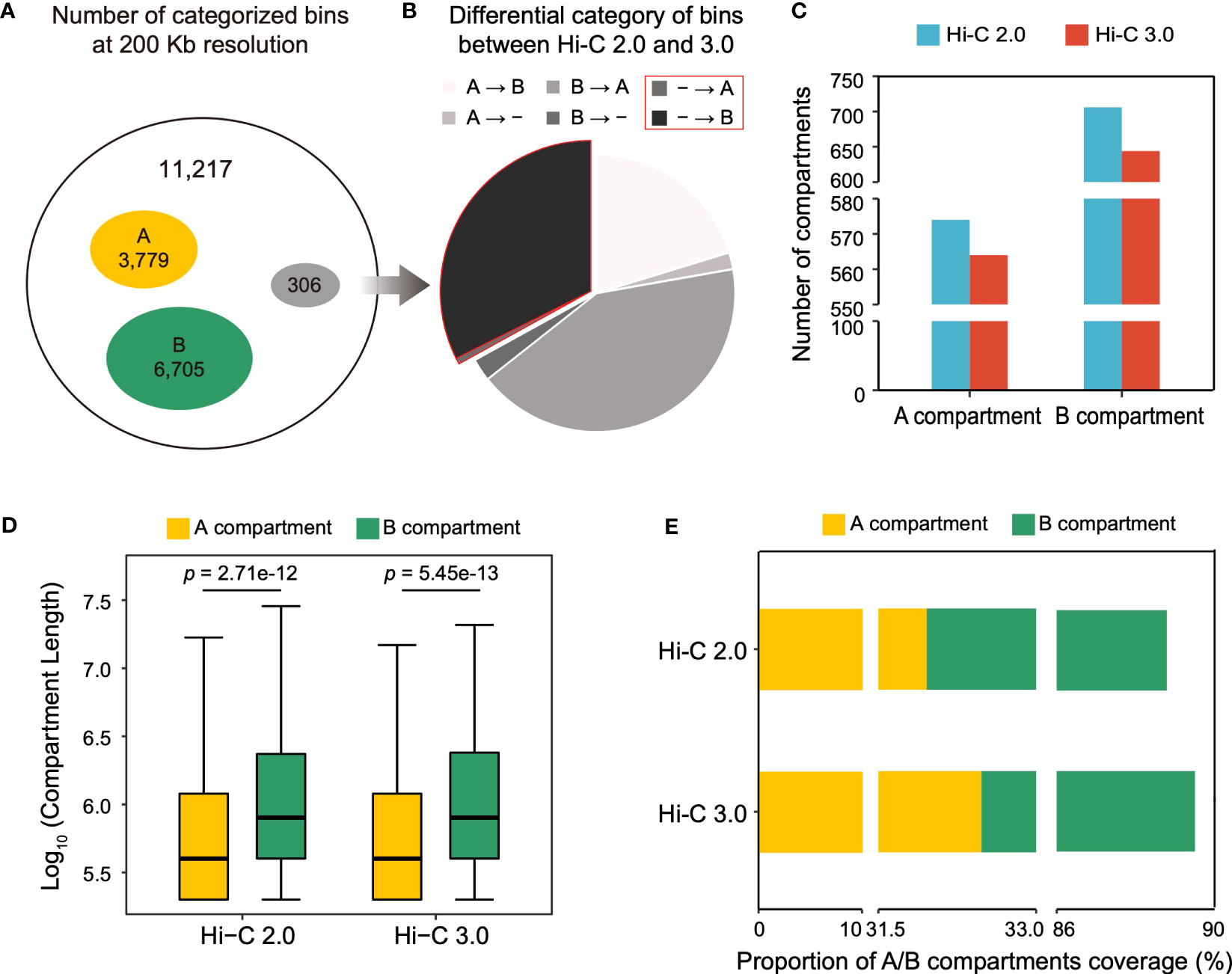
Frontiers An upgraded method of high-throughput chromosome conformation capture (Hi-C 3.0) in cotton (Gossypium spp.)

Mapping Nucleosome Resolution Chromosome Folding in Yeast by Micro

Nucleosome positions alone determine micro-domains in yeast chromosomes
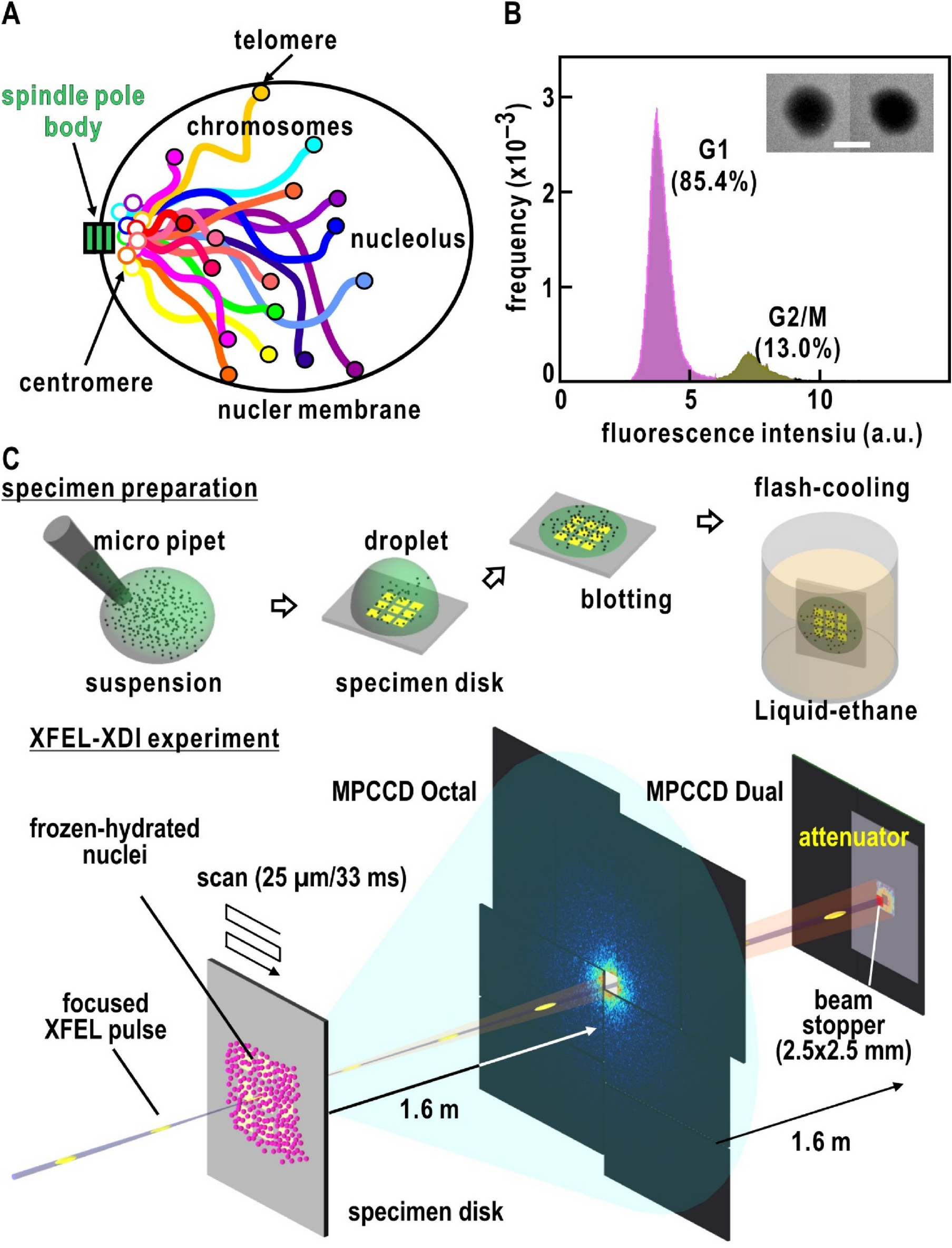
Ultrastructure and fractal property of chromosomes in close-to
Micro-C XL: assaying chromosome conformation from the nucleosome to the entire genome

Mapping Mammalian 3D Genome Interactions with Micro-C-XL
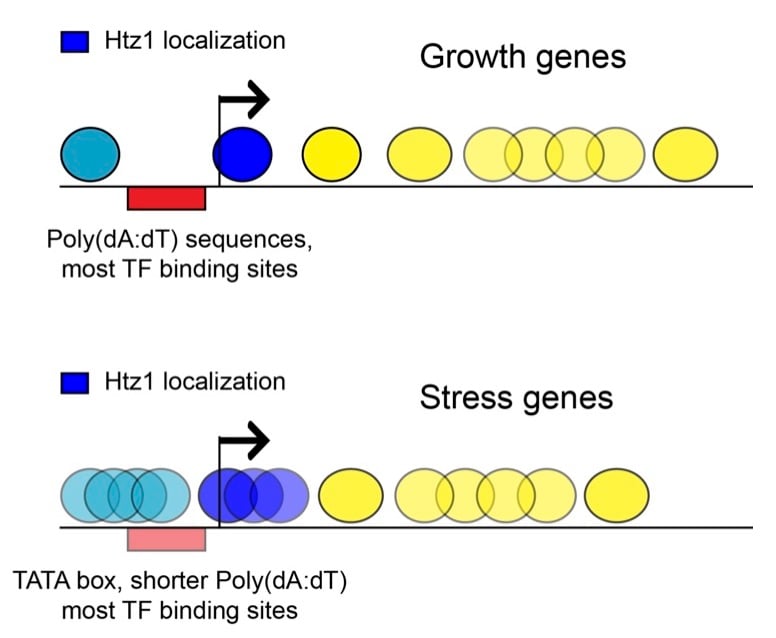
Chromatin Structure and Function
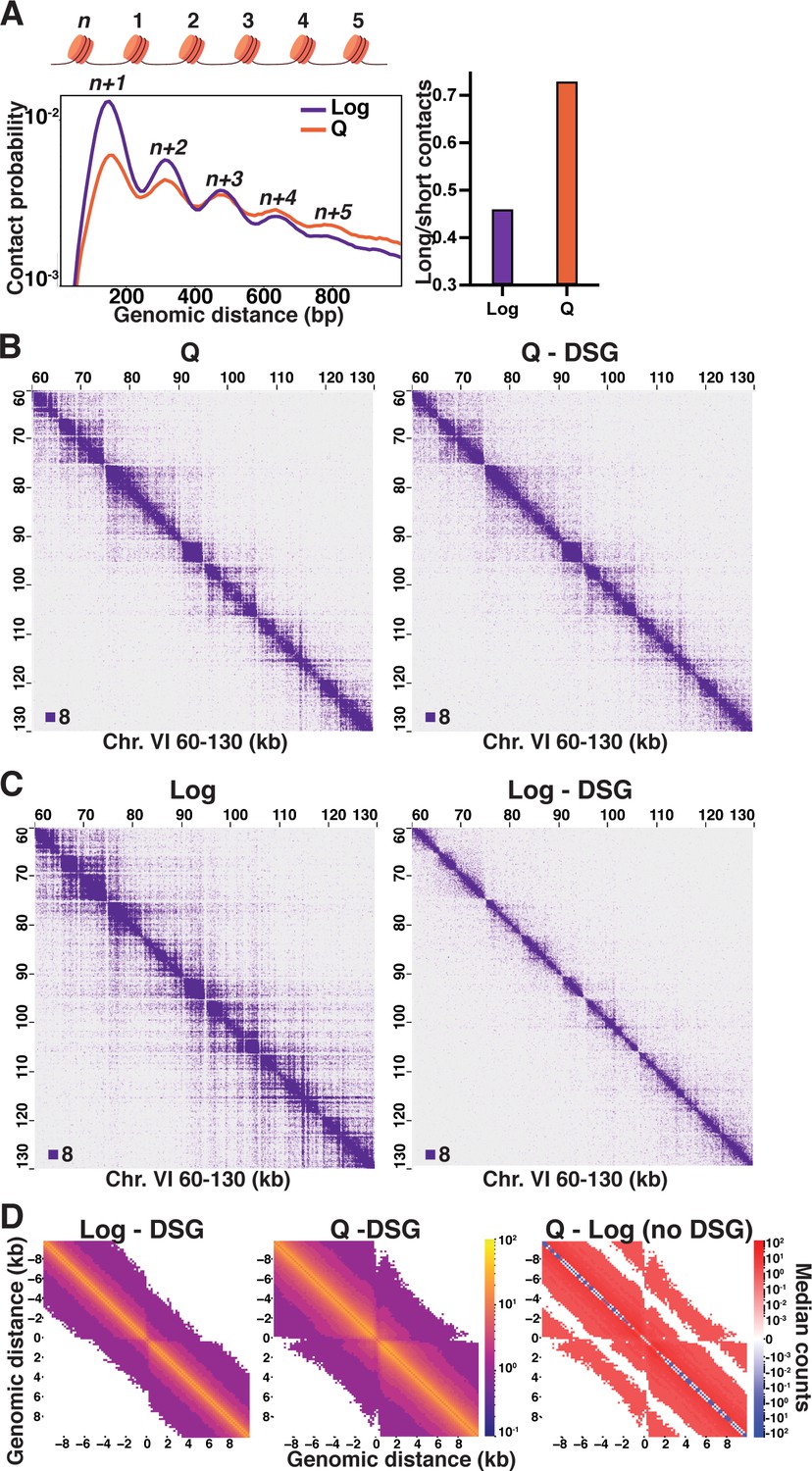
Local chromatin fiber folding represses transcription and loop extrusion in quiescent cells






